⇐ Previous topic|Next topic ⇒
Table of Contents
Power analysis
Summary
Before you do an experiment, you should perform a power analysis to estimate the number of observations you need to have a good chance of detecting the effect you're looking for.
Introduction
When you are designing an experiment, it is a good idea to estimate the sample size you'll need. This is especially true if you're proposing to do something painful to humans or other vertebrates, where it is particularly important to minimize the number of individuals (without making the sample size so small that the whole experiment is a waste of time and suffering), or if you're planning a very time-consuming or expensive experiment. Methods have been developed for many statistical tests to estimate the sample size needed to detect a particular effect, or to estimate the size of the effect that can be detected with a particular sample size.
In order to do a power analysis, you need to specify an effect size. This is the size of the difference between your null hypothesis and the alternative hypothesis that you hope to detect. For applied and clinical biological research, there may be a very definite effect size that you want to detect. For example, if you're testing a new dog shampoo, the marketing department at your company may tell you that producing the new shampoo would only be worthwhile if it made dogs' coats at least 25% shinier, on average. That would be your effect size, and you would use it when deciding how many dogs you would need to put through the canine reflectometer.
When doing basic biological research, you often don't know how big a difference you're looking for, and the temptation may be to just use the biggest sample size you can afford, or use a similar sample size to other research in your field. You should still do a power analysis before you do the experiment, just to get an idea of what kind of effects you could detect. For example, some anti-vaccination kooks have proposed that the U.S. government conduct a large study of unvaccinated and vaccinated children to see whether vaccines cause autism. It is not clear what effect size would be interesting: 10% more autism in one group? 50% more? twice as much? However, doing a power analysis shows that even if the study included every unvaccinated child in the United States aged 3 to 6, and an equal number of vaccinated children, there would have to be 25% more autism in one group in order to have a high chance of seeing a significant difference. A more plausible study, of 5,000 unvaccinated and 5,000 vaccinated children, would detect a significant difference with high power only if there were three times more autism in one group than the other. Because it is unlikely that there is such a big difference in autism between vaccinated and unvaccinated children, and because failing to find a relationship with such a study would not convince anti-vaccination kooks that there was no relationship (nothing would convince them there's no relationship—that's what makes them kooks), the power analysis tells you that such a large, expensive study would not be worthwhile.
Parameters
There are four or five numbers involved in a power analysis. You must choose the values for each one before you do the analysis. If you don't have a good reason for using a particular value, you can try different values and look at the effect on sample size.
Effect size
The effect size is the minimum deviation from the null hypothesis that you hope to detect. For example, if you are treating hens with something that you hope will change the sex ratio of their chicks, you might decide that the minimum change in the proportion of sexes that you're looking for is 10%. You would then say that your effect size is 10%. If you're testing something to make the hens lay more eggs, the effect size might be 2 eggs per month.
Occasionally, you'll have a good economic or clinical reason for choosing a particular effect size. If you're testing a chicken feed supplement that costs $1.50 per month, you're only interested in finding out whether it will produce more than $1.50 worth of extra eggs each month; knowing that a supplement produces an extra 0.1 egg a month is not useful information to you, and you don't need to design your experiment to find that out. But for most basic biological research, the effect size is just a nice round number that you pulled out of your butt. Let's say you're doing a power analysis for a study of a mutation in a promoter region, to see if it affects gene expression. How big a change in gene expression are you looking for: 10%? 20%? 50%? It's a pretty arbitrary number, but it will have a huge effect on the number of transgenic mice who will give their expensive little lives for your science. If you don't have a good reason to look for a particular effect size, you might as well admit that and draw a graph with sample size on the X-axis and effect size on the Y-axis. G*Power will do this for you.
Alpha
Alpha is the significance level of the test (the P value), the probability of rejecting the null hypothesis even though it is true (a false positive). The usual value is alpha=0.05. Some power calculators use the one-tailed alpha, which is confusing, since the two-tailed alpha is much more common. Be sure you know which you're using.
Beta or power
Beta, in a power analysis, is the probability of accepting the null hypothesis, even though it is false (a false negative), when the real difference is equal to the minimum effect size. The power of a test is the probability of rejecting the null hypothesis (getting a significant result) when the real difference is equal to the minimum effect size. Power is 1−beta. There is no clear consensus on the value to use, so this is another number you pull out of your butt; a power of 80% (equivalent to a beta of 20%) is probably the most common, while some people use 50% or 90%. The cost to you of a false negative should influence your choice of power; if you really, really want to be sure that you detect your effect size, you'll want to use a higher value for power (lower beta), which will result in a bigger sample size. Some power calculators ask you to enter beta, while others ask for power (1−beta); be very sure you understand which you need to use.
Standard deviation
For measurement variables, you also need an estimate of the standard deviation. As standard deviation gets bigger, it gets harder to detect a significant difference, so you'll need a bigger sample size. Your estimate of the standard deviation can come from pilot experiments or from similar experiments in the published literature. Your standard deviation once you do the experiment is unlikely to be exactly the same, so your experiment will actually be somewhat more or less powerful than you had predicted.
For nominal variables, the standard deviation is a simple function of the sample size, so you don't need to estimate it separately.
How it works
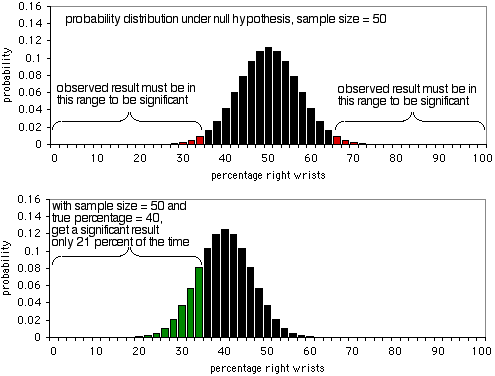
The details of a power analysis are different for different statistical tests, but the basic concepts are similar; here I'll use the exact binomial test as an example. Imagine that you are studying wrist fractures, and your null hypothesis is that half the people who break one wrist break their right wrist, and half break their left. You decide that the minimum effect size is 10%; if the percentage of people who break their right wrist is 60% or more, or 40% or less, you want to have a significant result from the exact binomial test. I have no idea why you picked 10%, but that's what you'll use. Alpha is 5%, as usual. You want power to be 90%, which means that if the percentage of broken right wrists really is 40% or 60%, you want a sample size that will yield a significant (P<0.05) result 90% of the time, and a non-significant result (which would be a false negative in this case) only 10% of the time.

The first graph shows the probability distribution under the null hypothesis, with a sample size of 50 individuals. If the null hypothesis is true, you'll see less than 36% or more than 64% of people breaking their right wrists (a false positive) about 5% of the time. As the second graph shows, if the true percentage is 40%, the sample data will be less than 36 or more than 64% only 21% of the time; you'd get a true positive only 21% of the time, and a false negative 79% of the time. Obviously, a sample size of 50 is too small for this experiment; it would only yield a significant result 21% of the time, even if there's a 40:60 ratio of broken right wrists to left wrists.

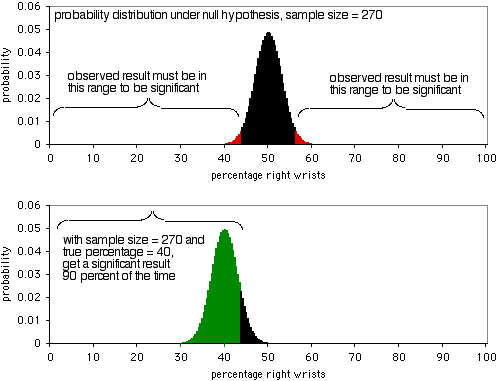
The next graph shows the probability distribution under the null hypothesis, with a sample size of 270 individuals. In order to be significant at the P<0.05 level, the observed result would have to be less than 43.7% or more than 56.3% of people breaking their right wrists. As the second graph shows, if the true percentage is 40%, the sample data will be this extreme 90% of the time. A sample size of 270 is pretty good for this experiment; it would yield a significant result 90% of the time if there's a 40:60 ratio of broken right wrists to left wrists. If the ratio of broken right to left wrists is further away from 50:50, you'll have an even higher probability of getting a significant result.
Examples
You plan to cross peas that are heterozygotes for Yellow/green pea color, where Yellow is dominant. The expected ratio in the offspring is 3 Yellow: 1 green. You want to know whether yellow peas are actually more or less fit, which might show up as a different proportion of yellow peas than expected. You arbitrarily decide that you want a sample size that will detect a significant (P<0.05) difference if there are 3% more or fewer yellow peas than expected, with a power of 90%. You will test the data using the exact binomial test of goodness-of-fit if the sample size is small enough, or a G–test of goodness-of-fit if the sample size is larger. The power analysis is the same for both tests.
Using G*Power as described for the exact test of goodness-of-fit, the result is that it would take 2109 pea plants if you want to get a significant (P<0.05) result 90% of the time, if the true proportion of yellow peas is 78%, and 2271 peas if the true proportion is 72% yellow. Since you'd be interested in a deviation in either direction, you use the larger number, 2271. That's a lot of peas, but you're reassured to see that it's not a ridiculous number. If you want to detect a difference of 0.1% between the expected and observed numbers of yellow peas, you can calculate that you'll need 1,970,142 peas; if that's what you need to detect, the sample size analysis tells you that you're going to have to include a pea-sorting robot in your budget.
The example data for the two-sample t–test shows that the average height in the 2 p.m. section of Biological Data Analysis was 66.6 inches and the average height in the 5 p.m. section was 64.6 inches, but the difference is not significant (P=0.207). You want to know how many students you'd have to sample to have an 80% chance of a difference this large being significant. Using G*Power as described on the two-sample t–test page, enter 2.0 for the difference in means. Using the STDEV function in Excel, calculate the standard deviation for each sample in the original data; it is 4.8 for sample 1 and 3.6 for sample 2. Enter 0.05 for alpha and 0.80 for power. The result is 72, meaning that if 5 p.m. students really were two inches shorter than 2 p.m. students, you'd need 72 students in each class to detect a significant difference 80% of the time, if the true difference really is 2.0 inches.
How to do power analyses
G*Power
G*Power is an excellent free program, available for Mac and Windows, that will do power analyses for a large variety of tests. I will explain how to use G*Power for power analyses for most of the tests in this handbook.
R
Salvatore Mangiafico's R Companion has sample R programs to do power analyses for many of the tests in this handbook; go to the page for the individual test and scroll to the bottom for the power analysis program.
SAS
SAS has a PROC POWER that you can use for power analyses. You enter the needed parameters (which vary depending on the test) and enter a period (which symbolizes missing data in SAS) for the parameter you're solving for (usually ntotal, the total sample size, or npergroup, the number of samples in each group). I find that G*Power is easier to use than SAS for this purpose, so I don't recommend using SAS for your power analyses.
⇐ Previous topic|Next topic ⇒
Table of Contents
This page was last revised July 20, 2015. Its address is http://www.biostathandbook.com/power.html. It may be cited as:
McDonald, J.H. 2014. Handbook of Biological Statistics (3rd ed.). Sparky House Publishing, Baltimore, Maryland. This web page contains the content of pages 40-44 in the printed version.
©2014 by John H. McDonald. You can probably do what you want with this content; see the permissions page for details.